July 10, 2018
A new generation of engineered vaccine candidates enters clinical testing
After decades of work, scientists are now advancing rationally designed vaccine candidates meant to induce a long sought-after broadly neutralizing antibody response.
Michael Dumiak
Armed with an understanding of HIV’s structure in minute detail and how antibodies bind to it, scientists are now ushering a new crop of engineered vaccine candidates into clinical trials, all in the hopes of eventually stimulating potent neutralizing antibodies that could block infection with the virus.
Vaccines work by training the immune system to respond to a specific pathogen before an infection ever occurs. The most crucial of these vaccine-induced immune responses are antibodies, typically Y-shaped proteins produced by activated B cells. Antibodies can latch on to and neutralize or inactivate viruses. For HIV, the focus has been on inducing broadly neutralizing antibodies (bNAbs): antibodies that can potently neutralize the many genetically different variants of HIV that are in circulation due to the virus’s unprecedented mutation rate.
Many scientists suspect bNAbs will be required to develop a highly effective, preventive HIV vaccine. Yet none of the vaccine candidates developed to date has been able to stimulate them. In fact, the antibodies elicited by current candidates can only neutralize a small fraction of HIV isolates that are typically transmitted from person to person (Nat. Med. 2018, doi:10.1038/s41591-018-0042-6).
This is not for lack of effort. Researchers have been exploring ways to develop bNAb-inducing vaccine candidates for decades. Now, thanks to significant scientific progress, these efforts are garnering broad interest and serious investment. About a dozen vaccine candidates designed specifically to initiate the induction of bNAbs are, or soon will be, tested in human volunteers.
These candidates come in different forms. Some are native-like trimers meant to resemble HIV’s outermost Envelope (Env) protein spike. Some others are computationally designed immunogens based on the precise epitopes on HIV Env that are targeted by bNAbs. There are also other strategies for invoking antibodies, as well as a slew of antibodies themselves being developed in an effort to protect against HIV infection.
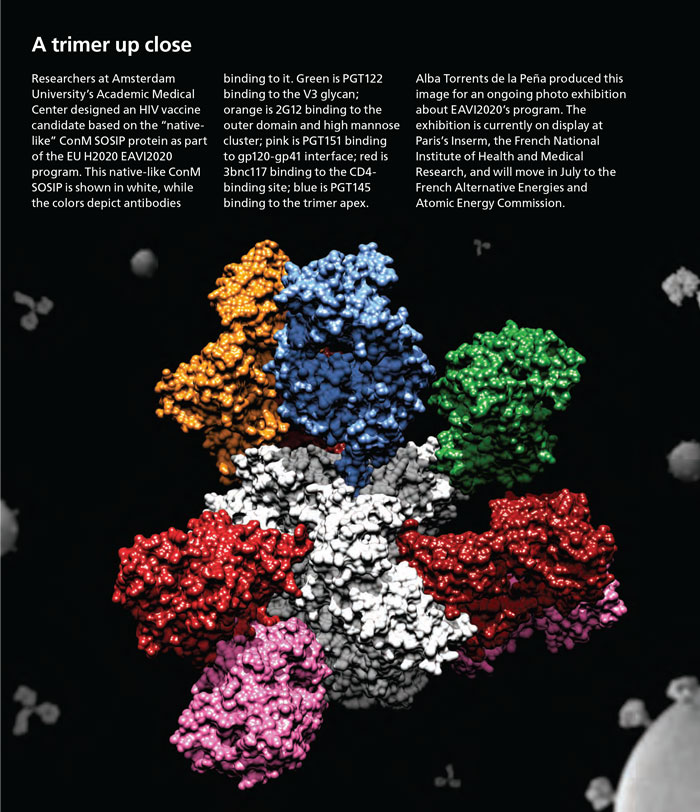 Image prepared by Alba Torrents de la Peña, PhD student in Rogier Sanders’ lab, in the department of medical microbiology at the Amsterdam University Medical Center
Image prepared by Alba Torrents de la Peña, PhD student in Rogier Sanders’ lab, in the department of medical microbiology at the Amsterdam University Medical Center
Awaiting results from efficacy trials
While many in the field are focused on advancing rationally designed, new-generation vaccine candidates, there is also a pair of ongoing efficacy trials that may illuminate additional paths to an HIV vaccine.
One is an efficacy trial in South Africa called HVTN 702, involving 5,400 volunteers. It is the follow-up to the RV144 trial of a canarypox viral vector prime and a gp120 Env protein boost. In the wake of the unexpected efficacy results from RV144, researchers extensively studied the immune responses the experimental vaccine regimen induced. While the vaccine seemed to produce antibodies, they were non-neutralizing. Instead they bound up HIV-infected cells until other parts of the immune system could come to the rescue, an action called antibody-dependent cellular cytotoxicity.
Researchers also reconfigured the candidates and dosing schedule to try to improve the efficacy of the regimen and to prolong the durability of the immune responses it induced. This included basing the candidates around clade C virus, which is most prevalent in South Africa. Results from HVTN 702 are expected by 2020. The trial is being supported by the US National Institutes of Health (NIH), the Bill & Melinda Gates Foundation, the South African Medical Research Council, the HIV Vaccine Trials Network, Sanofi Pasteur, GlaxoSmithKline, and the US Military HIV Research Program.
Another ongoing efficacy trial in South Africa is testing a so-called mosaic immunogen developed by Janssen and Dan Barouch,director of the Center for Virology and Vaccine Research at Beth Israel Deaconness Medical Center and a member of the Ragon Institute of Massachusetts General Hospital, the Massachusetts Institute of Technology, and Harvard University. It also carries with it an extensive support infrastructure, from Janssen, which is part of Johnson & Johnson, the Bill & Melinda Gates Foundation, and the NIH’s National Institute of Allergy and Infectious Diseases.
The mosaic antigen is computationally designed to provide maximum coverage against all currently circulating strains of HIV. The Imbokodo, or HVTN 705 trial, is enrolling 2,600 volunteers randomized to received either a four-valent cocktail expressing mosaic Env/Gag/Pol antigens boosted by a clade C gp140 soluble protein, or placebo.
Barouch says the studies leading up to Imbokodo show robust vaccine-induced antibody and T-cell responses: the antibody responses show functional activity, but not broad neutralization.
As for the slew of broadly neutralizing antibody approaches heading into the clinic, Barouch shows some skepticism. “While there are new strategies to trigger neutralizing antibody responses to particular epitope regions, it remains to be seen whether those strategies will be able to induce broadly neutralizing antibodies,” he says. “There are a lot of ideas, but not a lot of data, and we’re not even talking about human data. There’s very little animal data.”
“It’s a new era clinically of testing this antibody-based vaccine design concept,” says John Mascola, director of the National Institutes of Health’s Vaccine Research Center (VRC). “It’s certainly a very exciting time.”
 Electron microscopy reconstruction depicting antibodies attached to the HIV Envelope glycoprotein trimer at five sites of vulnerability shown in different colors. These five regions are the conserved epitopes that are currently being used by vaccine researchers as targets to design next-generation vaccine candidates. Image courtesy of Andrew Ward and Christina Corbaci at The Scripps Research Institute.
Electron microscopy reconstruction depicting antibodies attached to the HIV Envelope glycoprotein trimer at five sites of vulnerability shown in different colors. These five regions are the conserved epitopes that are currently being used by vaccine researchers as targets to design next-generation vaccine candidates. Image courtesy of Andrew Ward and Christina Corbaci at The Scripps Research Institute.
As remarkable as recent developments in the HIV vaccine field may be, none of these first-generation rational candidates is expected to induce bNAbs right off the bat. Researchers hope to use the early clinical trials to learn, Mascola says, not necessarily to solve. “We’re poised to learn a lot, even though the studies are what I would term as early stage. It’s still experimental medicine, meaning we need to learn from these studies to really understand the immune response to these vaccines.”
Vaccine development over the last century and a half was largely based on empiricism, or experimentation. Its guiding premise is that after conducting experiments in the lab and in animals, the only way to tell if a vaccine will work is to test it in humans. Many, if not most, vaccines were developed this way. But these trials are large and expensive, and in the 30 years that researchers have been trying to develop a vaccine against HIV, only one efficacy trial—the RV144 or so-called Thai trial—showed any efficacy at all. In this trial, the experimental vaccine regimen was 31 percent effective in preventing HIV infection.
“The history of vaccine experiments, and the vaccine experiments in this field, have not been stellar,” says John Moore, a professor of microbiology and immunology at the Cornell Weill Medical College. “Lots of immunogens over the years have tended to be put into trials because they exist. We need to understand what antigens are going to work best in animals, and at some point, humans. There are many aspects of these immunogens that remain to be understood.”
This is where the rational vaccine development effort comes in. It starts from hypotheses about what type of immune responses are thought to be protective, and then, working from that endpoint, researchers come up with vaccine components or antigens to try to trigger that response. Rational or hypothesis-driven vaccine design efforts are part of a larger vaccine movement against harder-to-combat pathogens for which an empirical approach would take too long and would be too expensive.
An advantage of this approach is that it doesn’t require going all the way to an efficacy study to determine if the vaccine candidate is having its desired effect. Researchers can learn this much earlier in the development cycle, allowing them to quickly improve upon the vaccine candidate and test it again, setting up an iterative process. Even though this new generation of rationally designed vaccine candidates will almost surely not deliver the final shot against the virus, researchers are poised to learn how to improve these candidates at a much faster pace than was possible in the past.
In the late 1990s, Moore and Rogier Sanders, now a virologist at the University of Amsterdam’s Academic Medical Center, began collaborating on what would be a decades-long effort. Early HIV vaccine trials testing subunits of the HIV Env which in the wild is a trimeric cluster of three glycoprotein (gp) 120 subunits and three gp41 subunits assembled into a spike—had not gone well.
The HIV Env spike is the only exposed viral protein, and therefore the target of all functional antibodies against the virus. But it is heavily glycosylated, providing multiple sugary decoys that shield the virus from the immune system. It is also notoriously unstable, constantly mutating and changing its shape to enable it to dock to and infect human cells (Immunol. Rev., 275, 161, 2017). Using monomeric Env subunits as vaccine antigens didn’t do the job. They induced plenty of antibodies, but they were not effectively neutralizing. Researchers suspected this was because these protein subunits were inadequate mimics of the native trimeric HIV Env protein (J. Infect. Dis., 173, 340, 1996; International Journal of Molecular Sciences, 19, 1241, 2018).
Creating a stable form of the Env trimer, the teams hypothesized, might yield a better immunogen. So the researchers engineered an SOS gp140 protein and manipulated it in the lab to try to stabilize it (J. Virol, 74, 627, 2000).
They introduced artificial mutations, including a disulfide bond between the gp120 and gp41 proteins, and made a genetic manipulation to try to prevent conformational changes that caused the trimer to fall apart. By doing so, Sanders and Moore and their colleagues were, by 2002, able to create a stabilized Env trimer. They called it SOSIP gp140 (J. Virol, 76, 8875, 2002).
But it turned out the protein wasn’t as stable as they’d hoped. “We realized the end product wasn’t really what we wanted it to be. It wasn’t stable enough and it wasn’t kept in the native conformation well enough, for long enough,” Sanders says.
The labs went on to work on other projects. Eight years would pass before they returned to the trimer stabilization efforts.
In 2006, a new idea emerged that would greatly influence antibody-based vaccine development. The Neutralizing Antibody Consortium, under the auspices of IAVI and led by Dennis Burton at The Scripps Research Institute, launched a research effort called Protocol G. The protocol’s aim was to collect samples from HIV-infected volunteers in sub-Saharan Africa, the US, UK, Australia, and Thailand, and screen them for antibodies with the unusual property of being neutralizing against a broad variety of viral strains in the laboratory.
Protocol G showed that a small subset of HIV-infected individuals is able to generate these antibodies over time. They are not able to provide much benefit to the infected person, as the virus can quickly mutate around this immune response, but researchers have long figured that it will be bNAbs such as these that, if induced by a vaccine, may be able to prevent HIV infection from ever occurring.
From Protocol G, scientists isolated an antibody called PG9, then another dubbed PG16, which were the first new bNAbs researchers had to work with in many years (Science, 326, 285, 2009). Mascola’s VRC soon after published on another broadly neutralizing antibody, VRC01 (Science, 329, 856, 2010).
The number of newly isolated bNAbs soon ran to a dozen, then dozens (Ann. Rev. Immunol., 34, 635, 2016). Some proved even better at neutralizing a broad swath of HIV isolates in lab tests. By genetically characterizing these antibodies, scientists began to understand precisely where these antibodies bind to HIV. This resulted in a map of antibody targets on the virus that could be exploited by vaccine researchers.
“The actual number of broadly neutralizing antibodies is pretty hard to characterize. But one way to look at it is from the major epitopes on the virus that the antibodies define. There are five major regions and maybe a sixth on the virus that are assigned by broadly neutralizing antibodies. And all of those sites are, in some fashion, a target of vaccine design,” Mascola says. These five epitopes comprise HIV’s CD4 binding site, the V1 to V2 apex, the V3 glycan super site, the membrane proximal region, and the gp120/gp41 interface region (see Image; Nat. Comms., 6, 8571, 2015).
With new antibody discovery continuing at an unprecedented pace, researchers returned to trimer stabilization efforts—this time with renewed funding, improved technologies, and a slew of antibodies in hand that allowed them to more easily attack the problem. “What really helped that effort was to have antibodies to the trimer, which allowed investigators to solve the crystal structure of the trimer, and therefore to better understand how to make it and how to stabilize it,” says Mascola. “It’s not a coincidence that this re-emerged along with the broadly neutralizing antibodies.”
In 2013, Moore, Sanders, and their teams were able to successfully stabilize HIV’s Env protein in an appropriately native-like conformation. They did this by deleting a string of 15 amino acids from their previous SOSIP construction and screening Env proteins from many isolates.
Moore, Sanders, and their colleagues in New York, Amsterdam, and La Jolla, California, stabilized an HIV gp140 protein called BG505 SOSIP.664 (PloS Path. doi.org/10.1371/journal.ppat.1003618). It was derived from a clade A virus isolated from a six-week-old Kenyan infant born in Nairobi, who had developed a bNAb response to HIV after being infected for two years.
Entering the antibody age
As vaccine researchers march new concepts into the clinic, more teams are also advancing multiple monoclonal antibodies for passive administration, which refers to direct injection or infusion of broadly neutralizing antibodies to try to treat, prevent, or possibly even cure HIV infection. This method avoids the need to stimulate the immune system to make these antibodies.
Several of these antibodies are approaching early-phase trials, while one antibody, VRC01, is already in an efficacy trial known as the Antibody Mediated Prevention (AMP) study (Current Opinion in Immun., 41, 39-46, 2016). VRC01 neutralizes an extensive panel of global HIV strains in the lab and has been shown to protect nonhuman primates against infection with SHIV, a monkey/human hybrid virus. The antibody has also proved to be safe and well tolerated in humans, and passive administration suppressed viral replication transiently in the VRC’s Phase I clinical trials VRC601 and VRC602.
A related antibody, VRC01LS, is in a Phase I safety trial evaluating the pharmacokinetics of the antibody in the serum and mucosa of healthy adults. This antibody differs only slightly from VRC01—it has a small amino-acid change designed to extend its half-life. The VRC is also working with Sanofi to advance a “trispecfic” antibody: an artificial antibody created with three “arms,” each from a different antibody (Science, 358, 85, 2017).
Several other antibodies have been in or will be entering Phase I trials shortly. Some of these include the VRC’s VRC07-523LS, 10E8VLS, and N6LS, and Rockefeller University’s 3BNC117 and 10-1074.
The Rockefeller antibodies have been through Phase I trials and have shown some suppression of viral rebound in HIV-infected volunteers, following interruption of antiretroviral treatment (Science, 352, 997, 2016). Now, they are being tested together in a Phase I trial, and researchers are also creating longer-lasting versions of these two antibodies. The antibody PGT121 is also slated for clinical trials, while many others are also in various stages of development.
Marina Caskey, an immunologist and associate professor of clinical investigation at Rockefeller University, is enthused by the many advances in developing vaccine immunogens. At least for now, she thinks the way ahead is clearer for passive administration.
“There’s proof of concept that it will work,” she says. But there is still much to learn about this approach as well. “We don’t yet understand the relationship with how much of an antibody you need in serum at the time of exposure in order to block infection from occurring,” she says. “We hope we will learn a lot about that in the AMP study.”
They then collected immunogenicity data in rabbits and nonhuman primates for the BG505 SOSIP.664 trimer, as well as another native-like trimer called B41 SOSIP.664, which was based on a clade B founder virus from an HIV-infected adult (Science, 349, 6244, 2015). The SOSIP trimers were each only able to induce neutralizing antibody responses against autologous virus, that is viruses from same strain as the sequence used to make the native-like SOSIP trimer. Even so, this was the first time an antigen showed an ability to provoke a strong and consistent neutralizing antibody response against an autologous virus with properties matching those of a transmitted virus. According to Sanders, it made an excellent starting point for iterative vaccine design.
“We’re trying to make immunogens that induce neutralizing antibodies against resistant viruses and to deal with the problem of diversity,” Moore says. “ If it were an easy thing to do, it would have been done a long time ago.”
Researchers then focused even more on how to optimize native-like trimers. The increasing use of cryo-electron microscopy and vastly more detailed modeling yielded unprecedented atomic-level resolution of HIV. “Those have been instrumental in designing further improved trimers, as well as imitations that made the technology applicable to envelope sequences from other viral strains,” Sanders says.
Now there are hundreds of these stable trimers, some of which are already in, or are being readied for, clinical trials. A vaccine candidate employing the BG505 SOSIP.664 gp140 protein as the antigen insert, with a GlaxoSmithKline adjuvant made up of a monophosphoryl lipid and the proprietary immune booster QS-21 combined in a liposome solution, will enter Phase I trials in the US and Kenya this summer.
Robin Shattock, head of mucosal infection and immunity within the department of medicine at Imperial College London, has long called for faster-moving, iterative, early-phase clinical studies, and that is what he is coordinating with the European AIDS Vaccine Initiative (EAVI 2020). The program is working to advance a total of seven SOSIP native-like trimer candidates into the clinic, as well as a single-chain construct based on a consensus sequence. Two stabilized SOSIPs are due to enter Phase I trials in early 2019, with another six candidates to follow later that year and in early 2020.
Seven of these eight candidates are coming from Sanders’s lab in Amsterdam, the other from Shattock’s (Immunol. Rev., 275, 161, 2017). “We’re looking at two different stabilization approaches to see which is the best,” Shattock says. “We’ve stabilized them molecularly, and we are taking a further step and stabilizing them chemically. We have made it almost like a rock and we want to know which of these gives us a flavor of antibody response that we have not seen before.”
As is often the case, HIV presents unique challenges, and the development of bNAbs is no exception. First of all they develop rarely—only in about 20 percent of infected individuals—and even then, only after years of exposure to the ever-mutating virus. Broadly neutralizing antibodies to HIV are also highly mutated as a result of undergoing multiple rounds of a process called somatic hypermutation. It is through this process that antibodies accrue the mutations that allow them to better bind to and neutralize HIV.
But highly mutated bNAbs are typically not able to bind to native HIV, which creates yet another challenge for vaccine scientists. The challenge is how to stimulate so-called germline antibodies that can bind to native HIV and then shepherd them through the mutation processes required to become bNAbs. Current thinking is that this may require vaccinating with a series of immunogens, each meant to facilitate the evolution of germline antibodies to those that are broadly neutralizing, and hopefully do so faster than what happens in natural infection.
Barton Haynes, director of the Duke Human Vaccine Institute in North Carolina and of the Duke Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery (CHAVI-ID), and colleagues are attempting to recreate the process of bNAb development by sequentially immunizing volunteers with four HIV gp120 Env proteins isolated from an HIV-infected individual who eventually developed bNAbs, along with the lipid adjuvant GLA-SE. A Phase I trial, known as HVTN 115, is currently enrolling volunteers to receive this multi-Env candidate that goes by the name of CH505 or EnvSeq-1. The first part of the study is meant to find the optimal dose of the first Env protein. In the second part of the study, researchers will administer the entire set of sequential Envs to see if they can kickstart the process of bNAb development in uninfected volunteers, and whether adding a DNA vaccine candidate, DNA Mosaic-Tre Env, can further improve the immune response.
“We’ve learned an awful lot about what happens, from an immunologic standpoint, when broadly neutralizing antibodies are made. We’ve also learned what happens when they are not made, when they are disfavored,” says Haynes. “Now we’re moving into a phase where we are combining the new structural knowledge we’ve gained with what has to happen immunologically.”
The Duke CHAVI-ID group is also running trials comparing CH505 produced from stable transfection with that produced by transient transfection to see if transient transfection, which is a faster method of manufacturing vaccine, delivers similar immune responses. “We’re trying to get to iterative Phase I trials,” Haynes says.
He and others are also developing a peptide liposome vaccine candidate based on the membrane-proximal external region (MPER) on HIV’s gp41 glycoprotein subunit. “It’s been very difficult to make, but we’ve now succeeded in making it and we’ve had a successful engineering run,” Haynes says. Toxicity studies are due to finish at the end of the year, and the researchers have an eye toward testing the MPER liposome in clinical trials in late 2019 or early 2020.
“A lot of these efforts are categorized as what Dennis Burton years ago called reverse vaccinology,” Mascola says. “The premise is, if you have an antibody that works pretty well, meaning it neutralizes the virus, you can design your vaccine to elicit that antibody.”
This is the goal of work by CHAVI-ID-backed researchers at The Scripps Research Institute in La Jolla. They, in partnership with IAVI, are advancing a vaccine candidate structurally designed to induce germline precursors to bNAbs. The Scripps group, including William Schief, Dennis Burton, Ian Wilson, and others, have employed computation-guided in-vitro screening to engineer a germline-targeting immunogen based on the outer domain region of HIV gp120.
This approach is referred to as structure-based vaccine design, and it is being applied to other vaccine development efforts as well, including against respiratory syncytial virus (RSV; Clinical and Vaccine Immunology, 23, 189, 2016, and 23, 243, 2016).
Scripps researchers took this “engineered outer domain,” or eOD, and figured out how to manufacture and deliver it in a cluster of multiple copies arranged on nanoparticles that are “self-assembling,” making the antigen look more like a virus. The resulting vaccine immunogen, eOD-GT8 60 mer, is now ready for Phase I trials. These trials will evaluate the ability of the candidate to expand the pool of B cells capable of making germline antibodies against the CD4 binding site.
The idea is that once germline antibodies are stimulated, perhaps researchers could then use the eOD candidate as the first in a sequence of immunogens, the others being more and more native-like synthetic trimers, to encourage the mutation and evolution of the induced antibodies to the point where they are broadly neutralizing.
Other structure-based vaccine candidates are also in development by researchers at the VRC. One candidate, FP-KLH, is based on the exposed epitope on the N-terminal region of HIV’s fusion peptide (FP) that is targeted by the antibody N123-VRC34.01. Recently published data show that immunizing with FP-KLH combined with a native-like trimer induced antibodies with promising neutralization breadth in mice, guinea pigs, and nonhuman primates (Nat. Med. 2018, doi:10.1038/s41591-018-0042-6). This work provides proof of principle for this epitope-based vaccine design and suggests the exposed N terminus of FP is a site of “exceptional HIV-1 vaccine promise,” the study’s authors conclude.
While the field largely rallies around this new crop of rationally designed vaccine candidates, there are still those that see the appeal of a more empirical approach. Burt Dorman, an 80-year-old biophysical chemist, argues in a recent profile by Adam Rogers in Wired that a classical, empirical approach to HIV vaccine development is still the best path forward. Dorman has also published, along with Haynes Sheppard at Global Solutions for Infectious Diseases, an essay calling for a systematic look at inactivated HIV vaccines (AIDS, 29, 125, 2015).
Anthony Fauci, head of the US National Institute of Allergy and Infectious Diseases, argues there is a role for both rational and empirical approaches in one of his commentaries (Science, 349, 386, 2015). “These approaches are coalescing into concomitant paths toward a safe and effective HIV vaccine.”
Michael Dumiak, based in Berlin, reports on global science, public health, and technology.